Scientists hijack cell machinery to destroy cancer-driving proteins
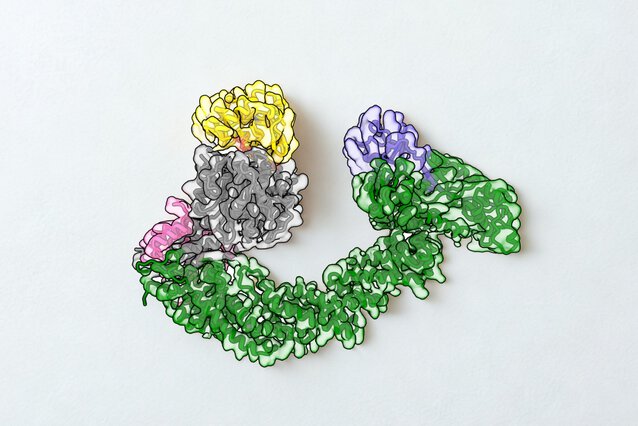

Every cell carries a built-in cleanup system to remove unwanted proteins, and researchers have learned to harness the enzymes in this system to destroy cancer-driving proteins. Scientists at the IMP and the University of North Carolina at Chapel Hill, have now captured this process in near-atomic 3D resolution, revealing its inner workings in unprecedented detail. Reporting in Nature Communications, they show that hijacked enzymes can destroy cancer-related proteins without compromising their normal function.
Healthy cells tightly regulate which proteins they contain and in what quantity, which allows them to respond to changing environmental conditions. To do so, they rely on specialised enzymes, called ubiquitin ligases, which label misfolded, damaged, or unwanted proteins with a molecular “destroy me” flag, triggering their breakdown by the cell’s waste disposal machine, the proteasome.
However, in cancerous cells, rogue proteins can slip through and accumulate, thereby contributing to disease progression. NSD2 (nuclear receptor-binding SET domain-containing 2) is one such protein, known to be a driver of several cancers, including multiple myeloma, acute lymphocytic leukaemia, and prostate cancer.
Ubiquitin ligases have inherent, natural targets, but scientists are now harnessing them to target other proteins for degradation. They design degrader molecules, which act as microscopic matchmakers that artificially pull the ubiquitin ligase and the target protein close together to form a three-way complex. The two main classes of such degraders are PROTACs – larger, two-ended molecules designed to bridge specific protein pairs – and molecular glues, smaller compounds that stabilise transient interactions between proteins. This orchestrated proximity facilitates the marking of the target protein and its subsequent degradation.
NSD2 has already been successfully flagged for removal using the ubiquitin ligase called FBXO22. Yet two key questions remained: exactly how does the degrader bring FBXO22 and NSD2 together, and does this interfere with FBXO22's normal function?
An international team of researchers at the IMP and at the University of North Carolina at Chapel Hill shed a new light on these issues. Their study, now published in the journal Nature Communications, solves the structure of the enzyme-degrader-target complex and shows that the right degrader molecule can maintain FBXO22’s natural function while allowing NSD2’s destruction.
Observing the matchmaking process in extraordinary detail
Using cryo-electron microscopy, the researchers reconstructed a detailed 3D image of the complex formed by the ubiquitin ligase, the degrader, and NSD2. It revealed that the degrader forms a bond with a specific site on FBXO22, locking it in place and positioning NSD2 so that it can be tagged for destruction. In addition, the structure showed that FBXO22 slightly shifts its shape to better expose this binding site and makes additional direct contacts with NSD2, strengthening the complex and making degradation more efficient.
“Seeing this complex at a very high resolution gives us a blueprint for how these degraders actually work,” says David Haselbach, head of the IMP’s cryo-EM Technology Platform. “It allows us to understand which interactions are essential and how we can improve them for future medical applications.”
A multitasking enzyme
One major concern with this therapeutic strategy is whether redirecting an enzyme like FBXO22 might interfere with its normal duties in the cell. Under natural conditions, FBXO22 already recognises and helps degrade its own set of target proteins. If a drug were to block this natural function, it could lead to undesired side effects.
The structural data showed that both natural and artificial target proteins attach to completely different regions of FBXO22. In other words, the degrader recruits NSD2 to a new site on the enzyme, leaving the original binding site for natural targets untouched.
Follow-up experiments confirmed that FBXO22 can form a larger complex containing both NSD2 and its natural substrate simultaneously, and cells treated with the degrader continued to regulate their normal targets. This suggests that the enzyme can “multi-task,” carrying out its natural role while eliminating the cancer-related protein.
“Our results provide an important proof of principle: it’s possible to design degraders that exploit cellular machinery without disrupting its normal function. This opens the door to more precise therapies that selectively remove harmful proteins while leaving healthy cellular processes intact,” says Haselbach.
Original publication
Kevin C. Robertson*, Sascha J. Amann*, Tongkun Liu*, Adam V. Funk, Xianxi Wang, Irina Grishkovskaya, Aamir Mehmood, John R. Tabor, Jacqueline L. Norris-Drouin, Cheryl H. Arrowsmith, Jon L. Collins, Yinglong Miao, Michael J. Emanuele, David Haselbach#, Lindsey I. James#, Nicholas G. Brown# “Structural basis of NSD2 degradation via targeted recruitment of SCF-FBXO22”. Nature Communications (2026). DOI: 10.1038/s41467-026-72235-9
* These authors contributed equally to the study.
# Corresponding authors
Further reading
The cryo-EM technology platform and David Haselbach’s research
How cells import their waste disposal machine into the nucleus