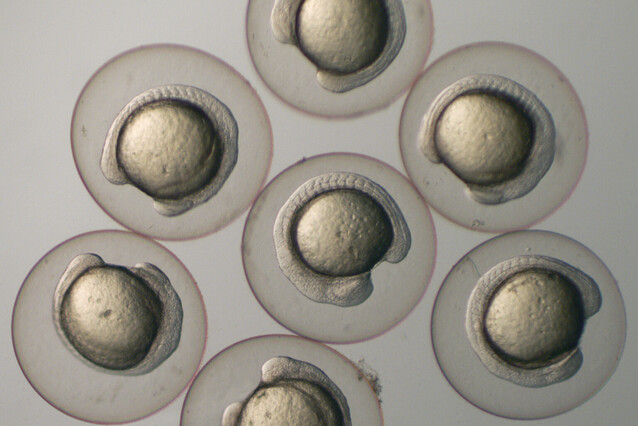
Finding out how embryos gain independence
After fertilisation, the nuclear genome of the embryo is mostly inactive, and maternal RNA and proteins in the egg perform cellular functions. During a process called maternal-to-zygotic transition (MZT), the embryo’s own transcriptional program is activated, and transcripts derived from the mother are cleared. A collaboration between the labs of Vienna BioCenter scientists Andrea Pauli and Stefan Ameres revealed detailed insights into the organisation of gene expression during this first major developmental transition in the life of vertebrates. Their findings have now been published in Cell Reports.
When a zygote, the first embryonic cell, forms after fertilisation, much of its protein and RNA content is pre-existing from the egg and guides the initial steps of cell division during the early stages of embryogenesis. During the MZT, the gene expression program is switched from the mother’s egg to the zygote, so that it produces all RNA transcripts necessary to carry out its cellular functions independently.
A deeper understanding of the transcriptional changes during this process has been hampered by technical limitations. Particularly, it has been difficult to distinguish between transcripts that are expressed by the zygote and those that have been deposited into the egg, a problem the scientists from the IMP, the Max Perutz Labs and Institute of Molecular Biotechnology of the Austrian Academy of Sciences have now solved, with their study published in Cell Reports.
The Pauli lab investigates the egg-embryo transition in zebrafish and has teamed up with the Ameres lab, which has long-standing expertise in RNA regulation and previously developed a technique called SLAMseq that allows to distinguish newly made transcripts from old ones. The scientists have now applied SLAMseq in a living organism, work that was driven by Vienna BioCenter PhD students Pooja Bhat and Luis E. Cabrera-Quio, to assess the transcripts during MZT in zebrafish and measure how their composition changes over time. “Through discussions in our RNA club at the Vienna BioCenter and in our RNA special research program, we realised that we could use the SLAMseq technology developed by my lab to understand how gene expression is coordinated during the MZT from a temporal perspective,” says Stefan Ameres.
“One remarkable finding of our study is that most transcripts present during gastrulation are still of maternal origin,” says Andrea Pauli. “When scientists talked about ‘zygotic transcription’, they referred to the small proportion of transcripts that are only made in the embryo – yet we show that most zygotic transcription occurs at genes whose transcripts are provided in the egg. These transcription events have been completely missed before.”
The team also found that different genes make the switch at different times and different speeds: genes that switch very quickly from maternal to zygotic expression include those that are involved in gene regulatory functions, most likely to ensure sustained gene expression during this critical developmental step.
The next steps in this research will be to determine how the switch from maternal to zygotic gene expression is implemented at the molecular level – and SLAMseq may very well be the technical solution to answer this question.
Original publication
Pooja Bhat, Luis E. Cabrera-Quoi, Veronika A. Herzog, Nina Fasching, Andrea Pauli#, Stefan L. Ameres#: “SLAMseq resolves the kinetics of maternal and zygotic gene expression during early zebrafish embryogenesis”. Cell Reports (2023), DOI: 10.1016/j.celrep.2023.112070.
# co-corresponding authors
Further reading
The dormant egg: scientists bring the ‘sleep mode’ of egg ribosomes to light