Scientists show how cells build protein destruction signals
In an international research effort, scientists have investigated how unwanted proteins in the cell are tagged for degradation. By combining state-of-the-art microscopy and sophisticated deep learning techniques, the team provides valuable insights into the choreography that governs protein regulation. The results of their study, published in the journal Nature Structural and Molecular Biology, shed light on previously unseen molecular processes.
Within the intricate molecular landscape inside of a cell, the orchestration of proteins demands precise control to avoid disease. While some proteins must be synthesised at specific times, others require timely breakdown and recycling. Protein degradation is a fundamental process that influences cellular activities such as the cell cycle, cell death, or immune response. At the core of this process lies the proteasome, a recycling hub in the cell. The proteasome degrades proteins if they carry a molecular tag formed by a chain of ubiquitin molecules. The task of attaching this tag falls to enzymes known as ubiquitin ligases.
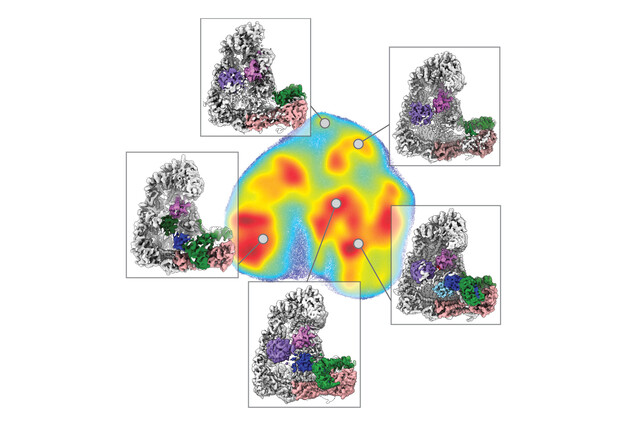
This process, known as polyubiquitination, has long been difficult to study due to its rapid and complex nature. To tackle this challenge, scientists at the IMP, the University of North Carolina, and collaborators employed a combination of techniques, integrating cryo-electron microscopy (cryo-EM) with cutting-edge deep learning algorithms. David Haselbach, a group leader at the IMP, explains: "Our aim was to capture polyubiquitination step by step through time-resolved cryo-EM studies. This method allowed us to visualise and dissect the intricate molecular interactions that take place during this process, like in a stop motion movie."
A biochemical timelapse
The study, now published in the journal Nature Structural and Molecular Biology, delves into the movements of the Anaphase-Promoting Complex/Cyclosome (APC/C), a ubiquitin ligase that drives the cell cycle. The mechanics behind APC/C’s attaching of a ubiquitin signal remained an unsolved puzzle. “We had a solid grasp of APC/C’s fundamental structure, a prerequisite for time-resolved cryo-EM,” says first author Tatyana Bodrug. “Now we have a much better understanding of its function, every step of the way.”
Ubiquitin ligases perform many tasks, including recruiting different substrates, interacting with other enzymes, and forming different types of ubiquitin signals. The scientists visualised interactions between ubiquitin-linked proteins and APC/C and its co-enzymes. They reconstructed the movements undergone by APC/C during polyubiquitination using a form of deep learning called neural networks. This was a first in protein degradation research.
The APC/C is a part of the large family of ubiquitin ligases (more than 600 members) that have yet to be characterised in this manner. Global efforts will keep pushing the boundaries of this field.
“A key to the success of our work was collaboration with several other teams. At Princeton University, Ellen Zhong’s software and programming contributions were key to uncovering new insights about the APC/C mechanism. Subsequent validation of these findings required the help of several other groups led by Drs Harrison, Steimel, Hahn, Emanuele, and Zhang,” highlights Nicholas Brown, Assistant Professor at the University of North Carolina. “Team effort was crucial to push our research over the finish line.”
The significance of this research extends beyond its immediate impact, paving the way for future explorations into the regulation of ligases, ultimately promising deeper insights into the mechanisms underpinning protein metabolism.
Original publication
Tatyana Bodrug, Kaeli A. Welsh, Derek L. Bolhuis, Ethan Paulаkonis, Raquel C. Martinez-Chacin, Bei Liu, Nicholas Pinkin, Thomas Bonacci, Liying Cui, Pengning Xu, Olivia Roscow, Irina Grishkovskaya, Michael J. Emanuele, Wei Zhang, Joseph S Harrison, Joshua P Steimel, Klaus M. Hahn, Ellen D. Zhong, David Haselbach*, Nicholas G. Brown*: “Time-Resolved Cryo-EM (TR-EM) Analysis of Substrate Polyubiquitination by the RING E3 Anaphase-Promoting Complex/Cyclosome (APC/C)” (2023). Nature Structural and Molecular Biology. DOI: 10.1038/s41594-023-01105-5
*co-corresponding authors
About the Vienna BioCenter PhD Program
Much of the work underlying this publication was done by a doctoral student of the Vienna BioCenter PhD Program. Are you interested in a world-class career in molecular biology? Find out more and apply in the current call until 15 October: https://training.vbc.ac.at/phd-program/