T cell differentiation and function
T cells are central cellular components of the vertebrate adaptive immune system that develop in the thymus from hematopoietic precursors. If rapidly acting innate immune defences fail to eradicate an invading pathogen, its molecular products trigger the expression of pro-inflammatory cytokines and co-stimulatory molecules and the mobilisation of specialised antigen-presenting cells (APCs) that carry pathogen-derived antigens to the secondary lymphoid organs to induce T cell activation. Activated T cells undergo rapid clonal expansion and acquire effector functions that allow migration to sites of infection, recruitment and stimulation of other immune cells, or direct killing of infected cells. Following pathogen clearance, the activated effector T cell pool contracts, leaving behind a clonally expanded population of long-lived memory T cells that protect the host from re-infection.
Depending on the context in which activation occurs, T cells differentiate into distinct types of effector cells with divergent functional characteristics tailored for protection against different types of microbial and abiotic challenges. Naïve CD4 T cells can differentiate into Th1, Th2, Th17, and TFH effector CD4 T cells whereas naïve CD8 T cells give rise to cytotoxic T lymphocytes. Additionally, chronic antigen stimulation of CD4 and CD8 T cells can result in a particular activation state known as dysfunction or exhaustion, while local tissue environments can invoke further distinctive features in non-lymphoid tissue-resident T cells. Concomitantly, a dedicated immunosuppressive subset of regulatory T (Treg) cells maintains self-tolerance and prevents lethal autoimmunity by dominantly suppressing the activity of autoreactive T cells. Our group is interested in understanding the molecular mechanisms underlying T cell differentiation and function in health and disease.
Epigenetic regulation of T cell activation states
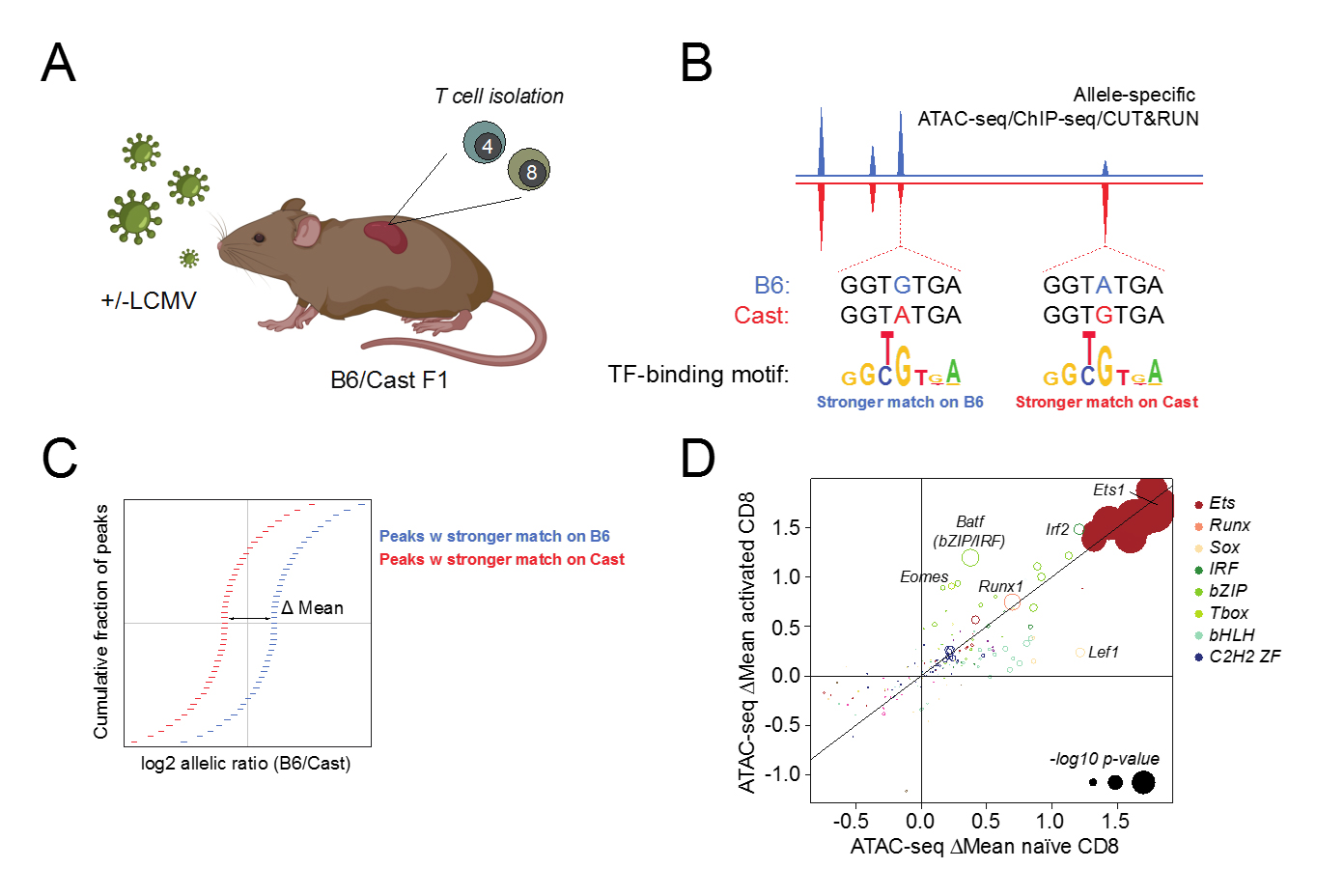
We have leveraged widespread polymorphisms in transcription factor binding motifs in wild-derived inbred mice to dissect mechanisms of gene regulation in T cells responding lymphocytic choriomeningitis virus (LCMV) infection. By analysing the effect of transcription factor binding motif variants on allele-specific chromatin accessibility in F1 offspring of C57BL/6 laboratory mice and Cast/EiJ wild-derived inbred mice, we have identified a number of TF families whose motifs most strongly affect chromatin accessibility, including Ets, Runx, and TCF/Lef (Fig. 1). Future studies are aimed at dissecting how these “heavy lifters” control chromatin accessibility, gene expression, and T cell function across different populations responding to acute versus chronic infection.
Foxp3-dependent regulators of T cell function
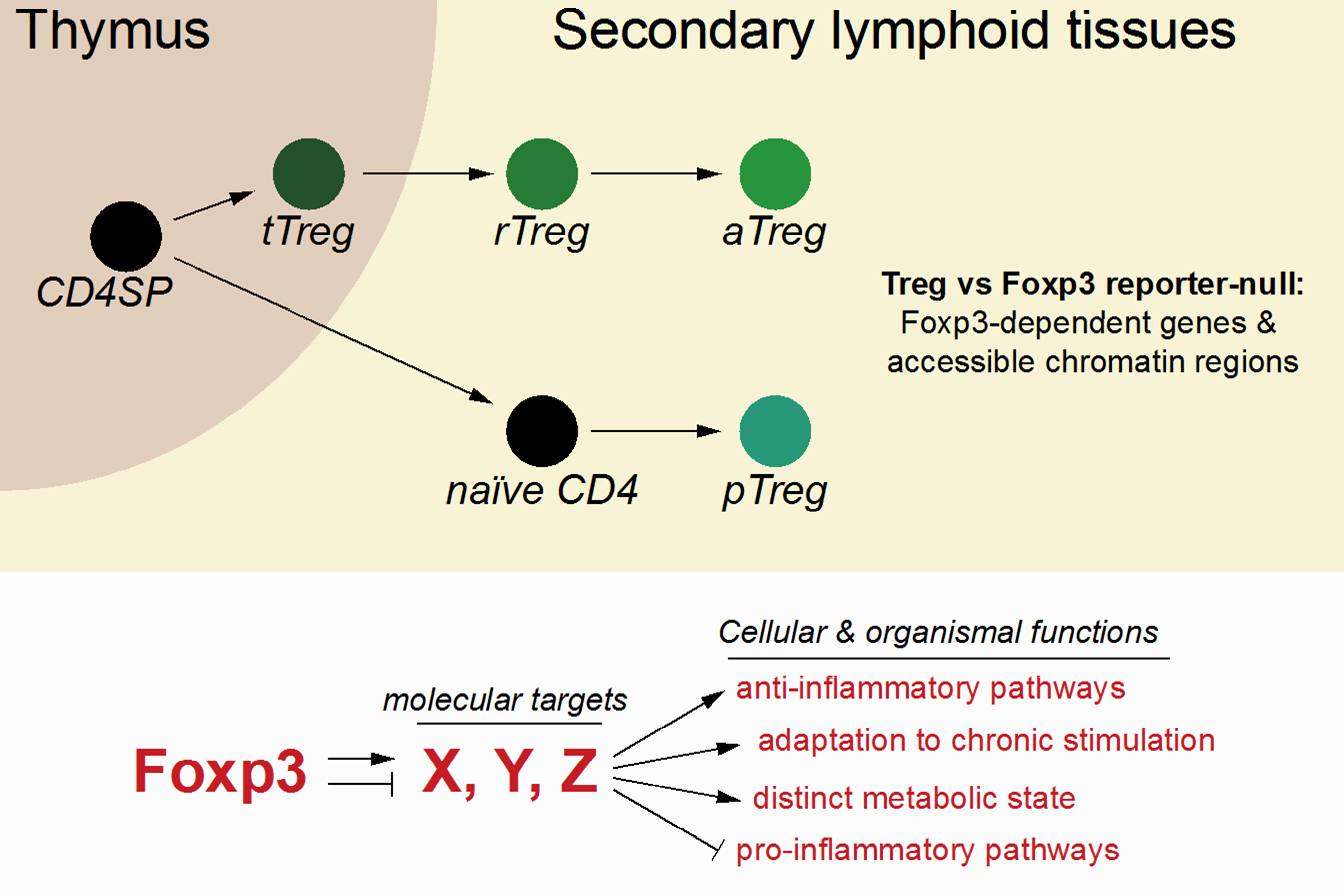
Regulatory T (Treg) cells are a dedicated population of immunosuppressive CD4 T cells that express the lineage-defining TF Foxp3 and dominantly suppress the activity of self-reactive effector T cells. Foxp3 expression is induced in a population of developing self-reactive thymocytes, supported by the activity of an intronic enhancer, CNS3. Treg cells and Foxp3 are critical for the prevention of lethal autoimmunity in mice and men and have been implicated in the regulation of tissue homeostasis and repair and the suppression of anti-tumour immune responses. How Foxp3 regulates its target genes to confer Treg cell identity is still unclear. Our work has shown that in contrast to major chromatin-remodelling transcription factors, Foxp3 does not affect the accessibility or expression of most of the genes to which it binds. Thus, Foxp3-dependent cellular functions likely depend on the regulation of just a small number of critical targets. Using a combination of genetic tools and genome-wide assays, we have identified Foxp3-dependent genes at various stages of Treg cell development and are currently investigating their functions.
Extrathymic differentiation of regulatory T cells
In addition to their development in the thymus, an independent pathway supports Treg cell differentiation from mature naïve CD4 T cells in secondary lymphoid tissues. These “extrathymic” or “peripherally-induced” Treg (pTreg) cells are thought to contribute to tolerance against food-derived antigens and the commensal microbiota; however, the mechanisms supporting their differentiation and their role in immune regulation remain largely unclear. We have developed genetic tools to study the development and function of pTreg cells and address the following questions: What are the conditions that support pTreg cell differentiation? How do Foxp3 and other transcription factors interact to define pTreg cell identity? What are the physiological functions of pTreg cells and how do these cells interact with other immune and non-immune cells?
Selected Publications
- van der Veeken J*#, Glasner A*, Zhong Y*, Hu W, Wang ZM, Bou-Puerto R, Charbonnier LM, Chatila TA, Leslie CS, Rudensky AY#. The transcription factor Foxp3 shapes regulatory T cell identity by tuning the activity of trans-acting intermediaries. Immunity (2020). *co-first authors. #co-corresponding authors
- van der Veeken J*, Zhong Y*, Sharma R, Dao P, Mazutis L, Pe’er D, Leslie CS, Rudensky AY. Natural genetic variation reveals key features of epigenetic and transcriptional memory in virus-specific CD8 T cells. Immunity (2019). *co-first authors.
- van der Veeken J, Gonzalez AJ, Cho H, Arvey A, Hemmers S, Leslie CS, Rudensky AY. Memory of inflammation in regulatory T cells. Cell (2016).
- Arvey A*, van der Veeken J*, Samstein R*, Feng Y, Stamatoyannopoulos JA, Rudensky AY. Inflammation-induced repression of chromatin bound by the transcription factor Foxp3 in regulatory T cells. Nature Immunology (2014). *co-first authors.
Join us
- Master students and Post-docs: Contact Joris van der Veeken with a letter of intent detailing why you want to join the lab.
- PhD students: Calls open 1 March and 1 September, apply here:
Vienna BioCenter PhD Program
Research questions and approaches
We use a YouTube plugin to display social media content on this page, which places requests to YouTube servers. These requests make your IP address visible to YouTube, who may use it in accordance with their data privacy policy. Please agree to make the YouTube video visible. You can find more information in our privacy settings.
Accept
