Advanced light microscopy and biophysics
How can light-based methods provide maximal information on biological processes? What are the limits to the information we can extract? How deep and fast can we look? How can we use extreme resolution to answer concrete questions? What answers can multimodal approaches deliver?
The development of novel optical methods like MINFLUX (see below) shows that the answers to these questions remain unexplored or incomplete, and that there is much to gain from synergistically combining expertise from diverse areas like optics, electronics, statistics, chemistry and biology. By solving transverse methodological challenges, we strive to push forward the maturation of light microscopy and profoundly influence life sciences along the way.
Super Resolution Microscopy
Fluorescence microscopy is an invaluable tool for exploring the structure and function of biological processes. It provides high specificity and contrast for the observation of cellular components (DNA, RNA, proteins, lipids, etc.) tagged with fluorescent molecules in a minimally invasive fashion, even allowing the study of live specimens. The spatial resolution of classical fluorescence microscopy is limited to hundreds of nanometers due to the diffraction of light; however, higher resolutions were unlocked with the development of the so-called super-resolution methodologies (stimulated emission depletion (STED) microscopy, photo-activated localization microscopy/stochastic optical reconstruction microscopy (PALM/STORM), among others)
In the last decade, achieving resolutions in the order of 10 nm to 100 nm became routine and has revealed details of subcellular organelles and new structures as well as ultrastructural anatomy in tissue, granting the Nobel Prize in Chemistry in 2014 to the developers of super resolution. Despite this revolution, the development of an ultimate microscope –revealing the precise nanometric location of all molecules of interest at all times without affecting the sample under study– remains elusive. Among several adversities, the photon budget of fluorescent probes is a fundamental bound for the trade-off between spatial and time resolution.
MINFLUX
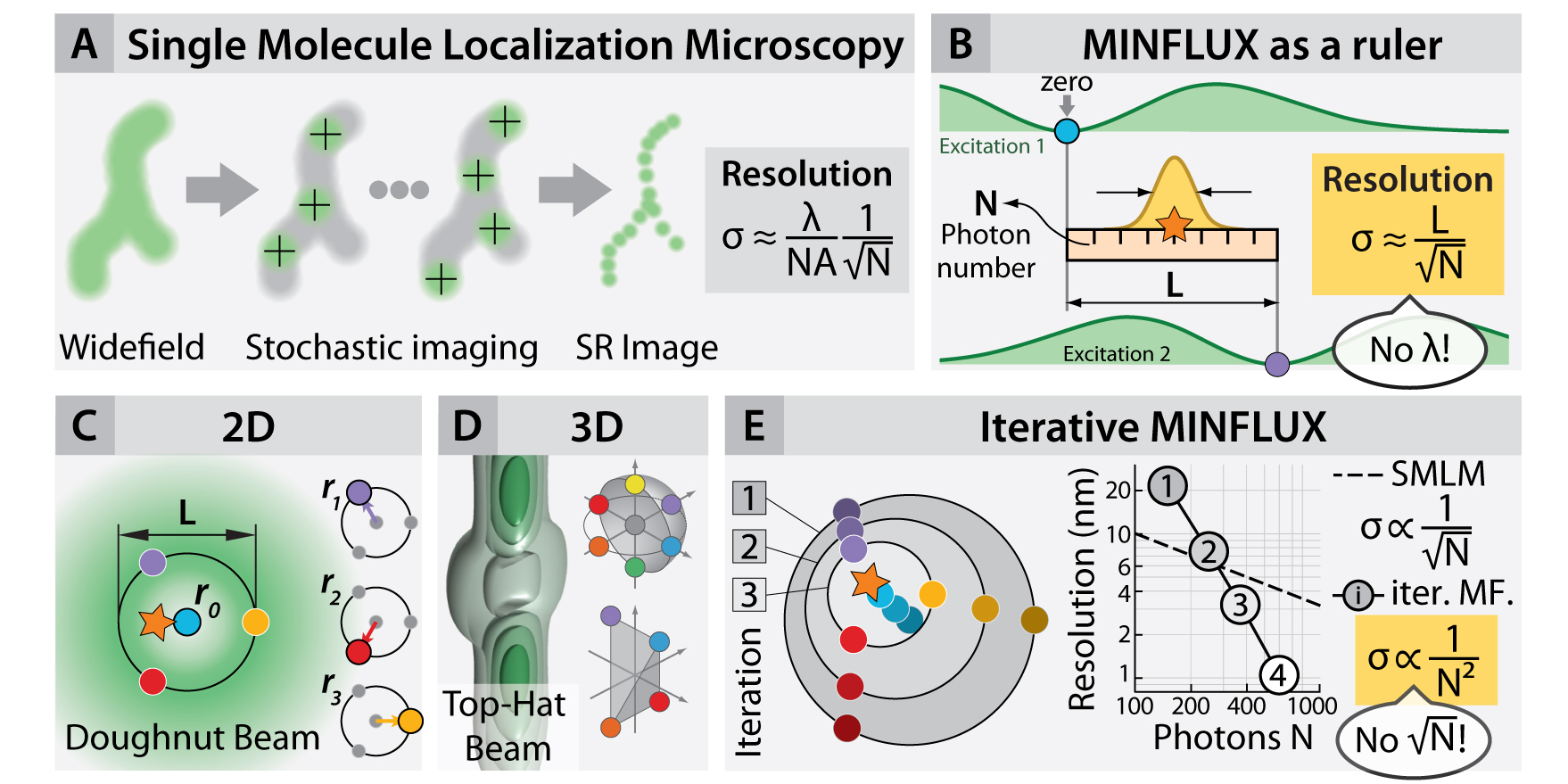
To overcome this, maximally informative luminescence excitation or MINFLUX, merges elements of information theory with (i) the single-emitter nature of PALM/STORM (fig. 1A) and (ii) the beam geometries typically used in STED (e.g the so-called “doughnut” beam). This technique (fig. 1 B–E) has shown that the information that each photon contains on the location of its emitter is a flexible quantity and that it can be dramatically increased in imaging and tracking applications (fig. 2B–D). Therefore, a given localization precision can be obtained by using much fewer photons (e.g. 20 times) than in conventional centroid-localization techniques, such as PALM and STORM.
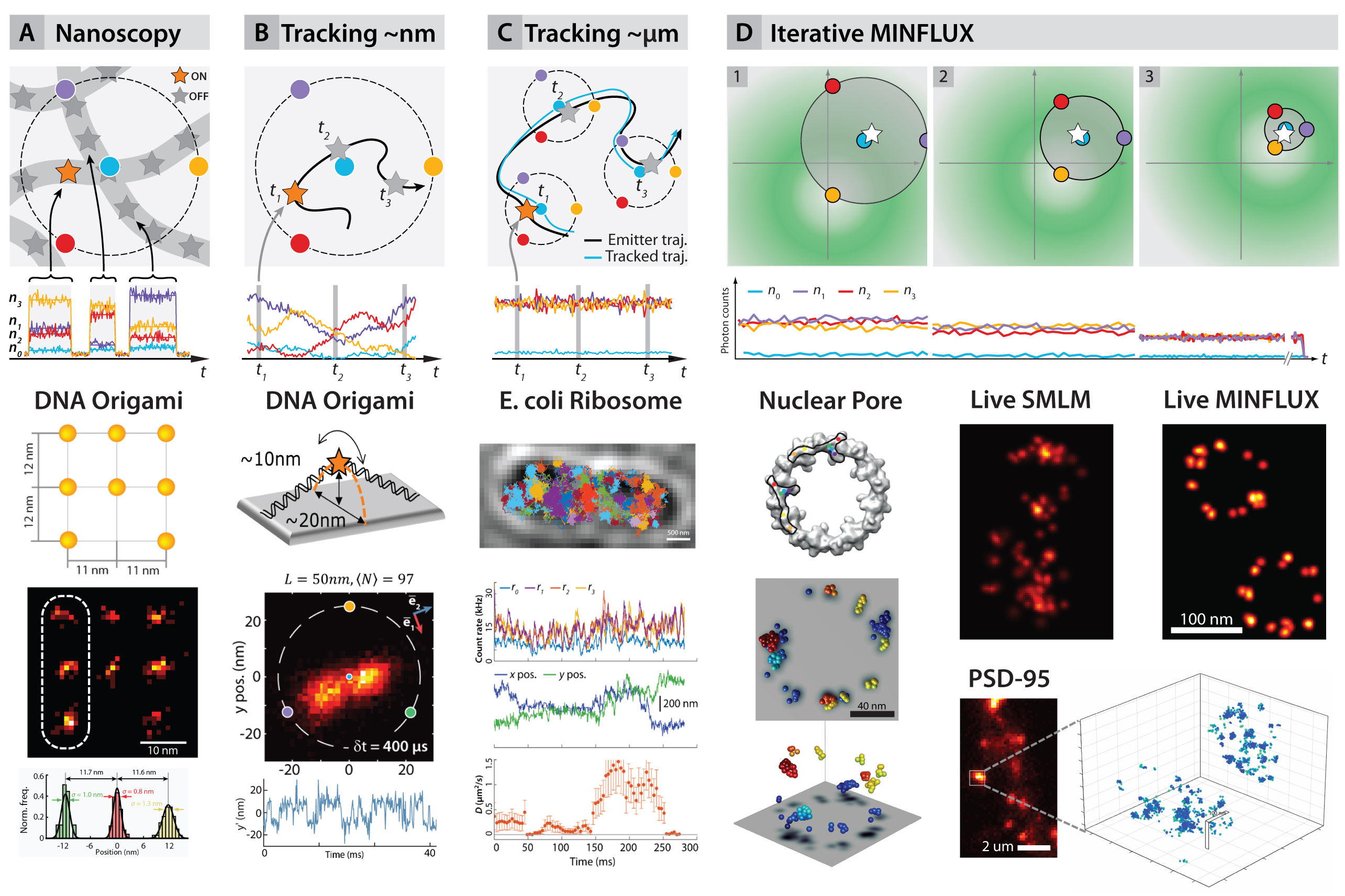
The concept was demonstrated in two and three dimensions by imaging DNA origami constructs and fixed and live cells, reaching 1– 3nm isotropic resolutions (fig. 2A,D). Tracking of the movement of DNA origami constructs (fig. 2B) resulted in ~2nm precision and sub ¬millisecond time resolution. Maximal photon economy was sought by tracking of 30S ribosomal subunit proteins in living E. coli fused with the photoconvertible protein mEOS2 (fig. 2C). Precisions of ~50 nm using 10 photons per localization were obtained, increasing the temporal resolution and the number of localisations per track hundredfold with respect to state-of-the-art tracking experiments under similar conditions.
Selected Publications
- Klaus C. Gwosch, Jasmin K. Pape, Francisco Balzarotti, Philipp Hoess, Jan Ellenberg, Jonas Ries & Stefan W. Hell. MINFLUX nanoscopy delivers 3D multicolor nanometer resolution in cells. Nature Methods 17, 217–224 (2020). doi.org/10.1038/s41592-019-0688-0
- Balzarotti, F. et al. Nanometer resolution imaging and tracking of fluorescent molecules with minimal photon fluxes. Science 355, 606–612 (2017). doi: 10.1126/science.aak9913
- Eilers, Y., Ta, H., Gwosch, K. C., Balzarotti, F. & Hell, S. W. MINFLUX monitors rapid molecular jumps with superior spatiotemporal resolution. Proceedings of the National Academy of Sciences of the United States of America 201801672 (2018). doi:10.1073/pnas.1801672115
- Gwosch, K. C. et al. MINFLUX nanoscopy delivers multicolor nanometer 3D-resolution in (living) cells. bioRxiv 734251 (2019). doi:10.1101/734251
Join us
- Master students and Post-docs: Contact Francisco Balzarotti with a letter of intent detailing why you want to join the lab.
- PhD students: Calls open 1 March and 1 September, apply here;
Vienna BioCenter PhD Program



